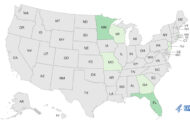
The tuna sushi Salmonella outbreak now includes 200 people from 21 states and the District of Columbia, to find out how those illnesses were related, scientists used DNA fingerprinting.
 Since 1996, PulseNet, a nationwide network of 87 laboratories, has worked together to identify and unravel foodborne illness outbreaks by posting the DNA fingerprints of pathogens that have caused confirmed cases of illness on a shared database.
Since 1996, PulseNet, a nationwide network of 87 laboratories, has worked together to identify and unravel foodborne illness outbreaks by posting the DNA fingerprints of pathogens that have caused confirmed cases of illness on a shared database.
“PulseNet has been a tremendously valuable tool for detecting outbreaks over the years,” said David Warshauer, deputy director of communicable disease at the Wisconsin State Laboratory of Hygiene, the lab that established the link between the tainted tuna and the outbreak patients. “For example, in this case we saw a cluster of Salmonella Bareilly and a cluster of Salmonella Nchanga. Once we see a cluster, we do a PGFE.”
PGFE is Pulsed Field Gel Electrophoresis, the method researchers use to determine the genetic fingerprint of strain involved. Once they have the print, they post it on the shared database. There are over 2,500 kinds of Salmonella, said Warshauer, but every subtype has its own fingerprint. When matching fingerprints start showing up on the database, it’s a good indication that there’s an outbreak.
Based on the PGFE results, health officials have been able to determine that the tuna sushi Salmonella outbreak now includes 190 people with Salmonella Bareilly infections and 10 people with Salmonella Nchanga infections. At least 28 people have been hospitalized and the investigation into the recalled tuna, produced by Moon Marine USA Corp., continues.